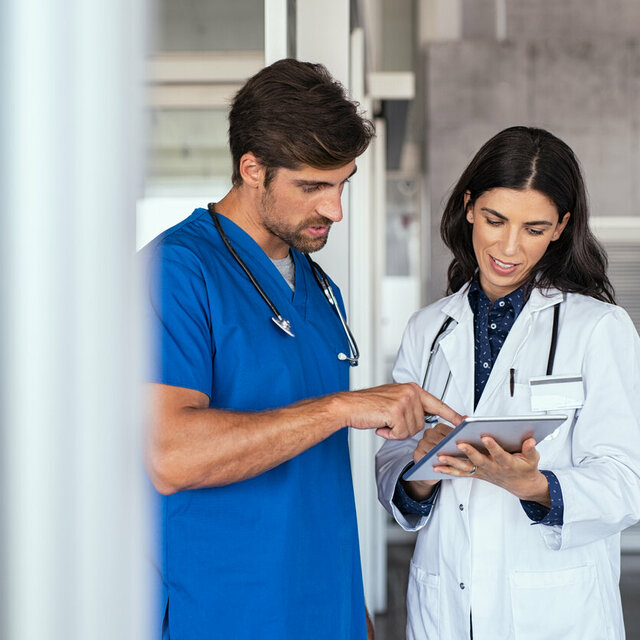
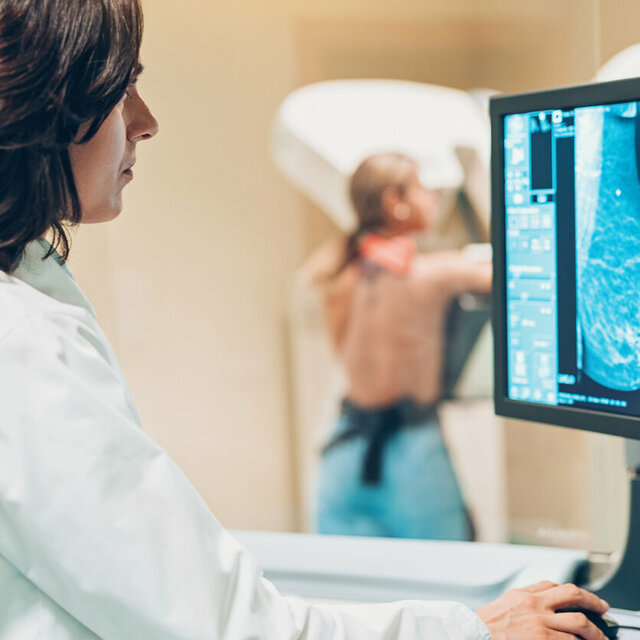
Expertise and experience from over 40 years of research
GBG Forschungs GmbH is one of the world's leading breast cancer research institutes. It conducts the German Breast Group’s clinical studies at an operational level.
As a basic principle, we are independent, academically oriented and impartial.
Our research focuses on studies that address academic questions based on concepts derived from the clinical activities of physicians. In contrast to studies initiated by pharmaceutical manufacturers, the approval of active substances is not the prime objective of our study concepts. We are an Academic Research Organization (ARO).


Development of innovative treatment options
More than 1,000 physicians at over 800 study centers are members of the GBG study group. Every year, more than 3,500 new patients participate in GBG studies. Clinical data from over 70,000 breast cancer patients have been collected so far.
Renowned breast cancer experts at the GBG are organized into subboards to develop innovative study concepts. New study protocols are developed and implemented together with breast cancer centers in Germany and around the world.
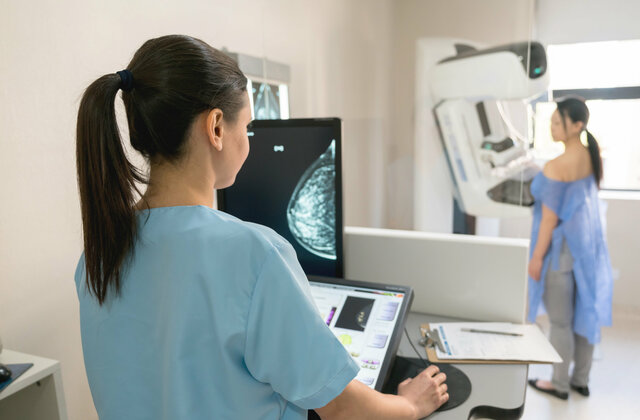
I've chosen GBG as an employer because it’s a fast-growing international company that offers professional development opportunities.
Develop state-of-the-art therapeutic approaches for breast cancer patients with us
![[Translate to English:] Eine Frau und ein Mann gehen einen Flur entlang.](/fileadmin/_processed_/a/d/csm_Stellenangebote_s_22b6f4d17c.jpg)
Questions to ask before participating in a clinical trial
1. What is a clinical study?
Clinical trials are carried out by internationally recognized experts at specialized breast cancer centers and clinics. The latest types of treatment and active substances are investigated in order to further improve the treatment of breast cancer patients. The aim is to find more and more individualized and effective therapies.
2. How do I benefit from taking part in a clinical study?
Patients with breast cancer who take part in a clinical trial receive innovative treatment options at an early stage that are not available in clinical practice yet.
It is a quality indicator if a breast center participates in clinical trials. By actively participating, you benefit from the scientific progress of treatment.
3. How can I participate ?
Every breast cancer has its own characteristics for which customized and targeted therapies are being developed. A number of trials are active at GBG that may be suitable for you. Find out more on the patient page.